Navigation
Achievements
IDA status to ICAR-NBAIM
Acquisition of the status of International Depository Authority (IDA) to “National Agriculturally Important Microbial Culture Collection” (NAIMCC) a unit of ICAR-NBAIM by World Intellectual Property Organization (WIPO), Geneva
National Agriculturally Important Microbial Culture Collection (NAIMCC), a unit of the ICAR-National Bureau of Agriculturally Important Microorganisms, Maunath Bhanjan, Uttar Pradesh, India under the aegis of Indian Council of Agriculture Research (ICAR) & Department of Agricultural Research and Education (DARE), Government of India, New Delhi has acquired the status International Depository Authority (IDA) by World Intellectual Property Organization (WIPO), Geneva, under Article 7 (1) of the Budapest treaty by its notification No. 338 w.e.f. July 28, 2020. Eighty two countries are part of this treaty and there are about 48 IDAs across 26 countries. The microbial resource centres having status of IDA mainly accepts and maintains microorganisms for patenting of work related to live organisms that have medical, agricultural and other uses. In view of this an agreement called as Budapest Treaty was passed in 1977 for deposition of microorganisms in culture collection centres for the purposes of patent procedure. NAIMCC will be the third IDA of the country after Microbial Type Culture Collection (MTCC), Chandigarh and National Centre for Microbial Resources (NCMR), Pune. As IDA, NAIMCC will be entrusted to conserve microorganisms used to develop patents and also to conserve newly described microbial taxa as a requirement for valid taxonomic publication. NAIMCC is a designated microbial repository for agriculturally important microorganisms (AIMs) under the National Biodiversity Act, 2002 and is a member of World Federation of Culture Collections (WFCC). Currently, NAIMCC holds accessions of 6907 AIMs including 2595 bacteria, 3981 fungi and 331 cyanobacteria.
The notification is available on http://www.ipindia.nic.in and on WIPO website with the link: https://www.wipo.int/treaties/en/ShowResults.jsp?country_id=ALL&start_year=ANY&end_year=ANY&treaty_all=ALL&search_what=N.
The Bureau wishes to share success story in detail. The correspondents may contact through email: director.nbaim@icar.gov.in or mobile 9650377776

Significant Research Work Done
- Repositories: A rich heritage for the future
The Bureau has established two repositories:
- National Agriculturally Important Microbial culture Collection (NAIMCC) with a total holding of 6800 live cultures of fungi, bacteria, actinomycetes and blue green alga (fig); and
- Microbial Genomic Resource Repository (MGRR) where in more than10000 accessions of genomic DNA, clones, plasmids, vectors and gene constructs are being maintained (www.mgrportal.org.in).
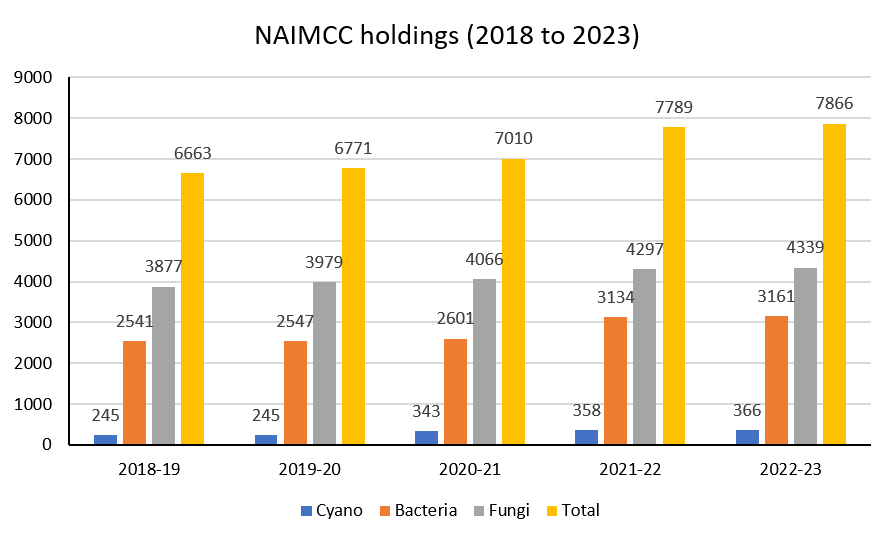
Fig: Annual Increment in Microbial Holdings
The Bureau is unique as it is the only Institution in the country mandated for preserving, conserving and utilizing microbial genetic resources for food and agriculture. The well characterized microorganisms are conserved for its sustainable use and are a wealth for national prosperity and posterity. Long-term Ex-situ preservation of microorganisms is carried through lyophilization of bacteria and sporulating fungi; and Cryopreservation of bacteria and fungi in liquid nitrogen (-1960C). Specific conservation protocols for recalcitrant microorganisms have been developed like gum based preservation of Chromobacterium violaceum. The recalcitrant microorganisms have also been conserved using modified lyophilization and step down freezing protocols of cryopreservators.
The holdings at NAIMCC includes 50 genera of bacteria and actinomycetes belonging to phyla Firmicutes, Proteobacteria and Actinobacteria and isolated from 29 states and an union territory Andaman & Nicobar Islands; 33 fungal genera particularly from class deuteromycetes, basidiomycetes and ascomycetes isolated from 23 states; and 18 genera of cyanobacteria belonging to orders Chroococales, Nostocales, Stigonematales and Oscillateriales isolated from 19 states. The Bureau maintains a core collection of agriculturally important microorganisms that have commercial importance and are used at present to develop formulations of biofertilizers, biopesticides and decomposers (fig).
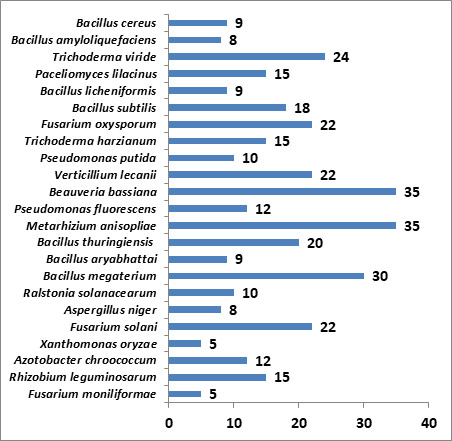
Fig: Core collection which includes frequently supplied cultures
The microbial germplasm maintained by the Bureau is being accessed by the academia for their research and academic activities while industries purchase the microbial accessions for development of commercial products to be used by farming community (Table)
Table: Supply of cultures, Safe Deposit and Revenue generated
| 2015-16 | 2016-17 | 2017-18 | 2018-19 | 2019-20 | Total | |
|---|---|---|---|---|---|---|
| Cultures supplied | 80 | 47 | 91 | 189 | 205 | 612 |
| Revenue (culture sale) | 0.62 | 1.11 | 3.52 | 0.98 | 3.95 | 10.20 |
| Safe Deposit | 0 | 0 | 13 | 30 | 8 | 51 |
| Revenue (Safe deposit) | 0 | 0 | 1.01 | 2.10 | 0.64 | 3.75 |
Total Revenue= Rs.13.95 lakhs
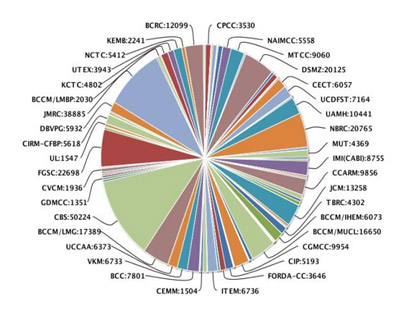
The name of species conserved along with its geographical data and characters are available at www.mgrportal.org.in and Microveda (an online database) and in form of published catalogues available on www.nbaim.org.in. NAIMCC is now in the global catalogues of Microorganisms
(Source http://gcm.wfcc.info/
StatisticgraphServlet)
ii. Bacterial diversity from different agroecological regions: Gold mine of novel microbes and genes
Extensive explorations throughout the country were carried out; culturable microbial diversity characterized for various traits and preserved under mid and long term conservation. The entire microbial community of certain sites has also been conserved in the form of 10 metagenomic libraries. The efforts to conserve the microbial diversity existing in wild have been further strengthened by initiating a project on ‘Indian soil Microbiome: wealth for national prosperity and posterity’ wherein concerted effort had been made to unlock the diversity of soil microorganisms of the entire country.

The efforts would provide base line information on abundance and diversity of microflora in different habitats. Such benchmarks are important to adjudge changes in soil microflora in the event of any calamity or even adoption of new management practices. First time, the Bureau has developed a database of Bacillus species from different extreme environments and identified niche specific bacilli. The most primitive group and the third domain of life, archaea that are difficult to culture have been isolated and preserved. The work has been carried out within the geographical boundary limits of the country and covers all the biodiversity hotspots in entire Indian sub-continent. About 20 different extreme environments including Bhitarkanika Mangroves, Sunderbans mangrove Manikaran, Vashist, Balarampur, and Bakreshwar thermal springs, Chilka lake, Orissa, Pulicat lake, Rann of Kutch, Gujarat, Eastern Uttar Pradesh, Leh cold desert, Rohtang pass, Kovalam district of Kerala, Jaisalmer, Rajasthan; Andaman & Nicobar island; Himalayan glaciers and Thar desert were explored. In addition four agroecological regions: Western Ghats and Coastal Plain; Eastern Ghats, Tamil Nadu Plateau and Deccan (Karnataka); Central Highlands (Malwas, Budelkhand, and Eastern Satpura); and North Eastern Hills (Purvanchal) have been explored. During these surveys more than 2000 microorganisms have been isolated, 238 have been identified, 6 metagenomes were analysed and 40 microorganisms of rare occurrence were preserved. Some rare cyanobacteria namely Chamaesiphon sp. SB2, Leptolyngbya antartica SK2, Hapalosiphon sp. BG2, Trichormus azollae SB1 Hapalosiphon sp. SB10 Chroococcidiopsis cubana HC1 Pleurocapsa sp HC3 Chroococcus sp. SB4 were isolated from extreme environments (Atri hot springs, Brahamagiri and Bhiterkanika, Odisha and Leh). The microbial diversity of Valley of Flowers National Park – a UNESCO national heritage site was deciphered. The population density of ~2.0 X 109 cfu/g was recorded. The cultivable bacterial diversity was active for production of hydrolytic enzymes like cellulase (21%), amylase (34%), β-glucosidase (35%), laccase (25%) and was also endowed with PGP like production of ammonia (68%) and siderophore (41%), solubilization of Zn (25%) and K (11%).
- Amplification of full length genes for Ectoine synthesis ectABCD from Halomonas and Virgibacillus ; Glycine Betaine genes gbsAB from Bacillus subtilis and ABC transporter OpuA – OpuAA, OpuAB and OpuAC from halotolerant Bacillus isolates was carried out.
- Developed Indicator microorganisms for
- Soil health: site Modipuram stn long term fertility experiments
There was high value of alpha diversity index at the organically managed soil whereas the inorganically managed soil was having the lowest value which shows that there is a loss of biodiversity at the inorganically managed soil. Enzyme assays also showed maximum activities in organically managed soil as compared to inorganically managed soil.The Linear discriminate analysis was performed to measure the most affected genera under various management practices. It was found that Gemmatimonas was the most significantly affected genera under organically managed soil microbiome.
- Pesticide contamination: Site Umari Village, Chinhat, Lucknow
There was significant loss in the representation of members of phylum Acidobacteria, Actinobacteria and Gemmatimonadetes at contaminated site. The Linear discriminate analysis showed that Marinimicrobium was the significantly affected genera due to lindane contamination. The heat map depicts that at the lindane contaminated site there was high abundance of PAH degrading genera like Pseudomonas , Sphingomonas , Bacteriodia etc.
- Arsenic contamination: Balia district
Comparative taxonomic and functional analysis between different samples revealed changes in relative abundance of bacterial community due to As contamination. At genus level in As contaminated soils, representative hits of Providencia genus were low in comparison to control. Genera Anaerolinea, Nocardiopsis, Pyrinomonas, Serratia, Streptomyces and unclassified Gemmatimonadetes gave significantly higher number of hits in As contaminated soil than control.
- Description of two novel bacterial species
- ICAR- NBAIM described a novel species of moderately haloalkalophilic actinobacteria isolated from salt crusts of Panamik hot spring, Leh, J&K and the nomenclature proposed is Nesterenkonia icaraensis sp. nov.
- A moderately halophilic, Gram negative, aerobic bacterium belonging to the genus Halomonas was isolated from soil of Pentha beach, Odisha. Based on the phenotypic, chemotaxonomic and phylogenetic characteristics, strain D1-1T represents a novel species in the genus Halomonas for which the name Halomonas icarensis sp. nov. have been proposed.
- Microbial Genomicss
Bureau has been continuously involved in deciphering capabilities of microbes by genomic analysis tools and techniques. With countries first complete draft genome sequencing of Mesorhizobium ciceri strainCa181 in 2011, whole genome sequencing of various agriculturally important microbes Exiguobacterium profundum PHM 11, Bacillus subtilis RC 25, Pseudomonas azotoformans SC 14, Chromohalobacter salexigens ANJ 207, Fusarium udum F-02845, Pseudomonas koreensis P2, Staphylococcus xylosusLSR_02N, Brevibacillus borstelensis LCHU R05 have been done and information on stress responsive traits, metabolic pathways and transporter profiles have been obtained.
Besides this environmental shotgun sequencing of various niches like Khardoong La Subglacial Soil, Pangong Lake, Leh, Mangrove soil, Saline soil, Solid waste, Wheat rhizosphere from different regions as well as saline soil, Arsenic contaminated soil; Valley of Flowers and Termite gut have been done and information on composition and functions have been derived from multitudes of uncultivable prokaryotic and eukaryotic species and complex microbial population existing in particular environment. Bureau has also taken lead in standardizing protocol for recovering all proteins recovered directly from environmental samples at a given time i.e. metaproteome and first time reference set of proteomic data of microbial communities associated with maize rhizosphere have been made available. Subsequently metaproteome of wheat rhizosphere from saline and non saline region has also been compared.
Reference
Whole Genome Sequence Submitted In NCBI
MRSV00000000; MZZI00000000; MZZJ00000000; MZZK00000000; NIFK00000000; NIGO00000000;
Bioproject ID: PRJNA271204Biosample ID: SAMN03273297; Bioproject ID: PRJNA271205 under accession number JXAV00000000; BioProject ID: PRJNA271205; Biosample ID: SAMN03273298; Bioproject ID: PRJNA40923 Biosample ID: SAMN02470606 Accessession Number: ASTL0000000.1
Metagenome submissions
At NCBI
- BioProject ID:PRJNA192218 Biosample: SAMN01940375;
- BioProject ID: PRJNA206543, Biosample ID: SAMN02401017
- BioProject ID:PRJNA382608;Biosample: SRP103712
- BioProject ID:PRJNA382604; Biosample: SRP103718
- BioProject ID:PRJNA382603; Biosample: SRP103716
At JGI GOLD Genome online database
- Gold study id:Gs0136013; Biosample (GOLD Biosample ID: Gb0196815; Biosample ID: Gb0196814; Biosample ID: Gb0196813
- Gold study id:Gs0132850; Biosample (GOLD Biosample ID: Gb0159122;
At MG RAST
- Metagenome MG-RAST IDs: mgm4821297.3 (C1_R.fastq) ; mgm4821985.3 (D1_R.fastq); mgm4821996.3 (H1_R.fastq)
At European Nucleotide Archives (ENA) of EBI
Study : PRJEB30074; Secondary Accession : ERP112485
Metaproteome data submitted at ProteomeXchange
- The mass spectrometry proteomics data in the ProteomeXchange Consortium via the PRIDE partner repository with the dataset identifiers PXD014519; PXD015387
- Diagnostic markers for fungal pathogens
- Fusarium

Development of PCR based diagnostic markers for Fusarium based on gene sequences for exopolygalacturonase (Pgx), intergenic spacer (IGS) and alcohol dehydrogenase (ADH)
2. Alternaria
A new diagnostic marker based on noxB gene has been developed and validated for Alternaria spp. on nine different crop species.

3. Rhizoctonia solani
A colorimetric LAMP assay has been standardized for detection of Rhizoctonia solani based on AG-1 Polygalacturonase gene.

VII . Composting through rapid decomposition of lignocellulosic biomass
A consortium of four hyper lignocellulolytic fungi (Trichoderma viride, Aspergillus niger, Pleurotus florida and Phanerochaete chrysosporium) recommended for composting of lignocellulosic waste. The consortium has been used for composting of diverse agricultural wastes such as paddy and wheat straw, soybean trash, pearl millet, and maize residues effectively. Mature compost can be obtained within 65-70 days by pit or windrow methods.
The technology involves a product in the form of microbial consortium. A carrier based inoculum is available in pack of 1kg and is sufficient for conversion of 1 ton of biomass. Mature compost can be obtained within 70-75 days by pit or windrow methods. By this process N (1-1.5%) and P (0.3-0.5%) enriched compost can be prepared from diverse crop residues within 70-75 days. The application of nutrient enriched compost in soil leads to a significant increase in the soil fertility status, in terms of microbial biomass, N and available P, enhancing the overall physical, chemical and biological health of soil. The supply of 120 kg N, 40 kg P2O5 and 30 kg K2O through application of 10 t compost is an environmental friendly option which can reduce the inputs of chemical fertilizers which are polluting our environment.
VIII. Microbe based technologies for nutrient management
The agriculturally important microorganisms (AIMs) obtained under various projects related to diversity are characterized and maintained at the Bureau under the umbrella of NAIMCC serving as bank for effective microorganisms which can be used for diverse agricultural applications like in biofertilizers, biocontrol agents and inoculum for fast composting technologies. The Bureau has made significant progress in this direction and has developed bioformulations to be used as biofertilizers, biocontrol agents, abiotic stress alleviators, agricultural residue decomposers etc.
1. Psychrotolerant P solubilising bacteria for enhanced growth of crop plants

Four psychrotolerant and plant growth promoting bacteria (PGPB) were isolated from cold deserts of the Arunachal Pradesh and identified as Pseudomonas arsenicoxydans P1; Pseudomonas koreensis P2, Pseudomonas koreensis P3 and Paenibacillus dendritiformis P4 based on 16S rRNA gene sequencing. Under green house conditions, all the strains remarkably enhanced root and shoot dry weight in chickpea, maize and wheat (fig).
Bio Phos and Bio Phos+ are liquid formulations of P- solubilizing bacteria containing Paenibacillus tylopili and Kluyvera cryocrescens respectively. Phosphorus solubilising bacteria play crucial role in dissolution or insoluble source of phosphorus through secretion of organic acids and other metabolites. Inoculation of P solubilizer helps to augment 15 to 20 kg P2O5 ha-1.
2. Assessment of Sulfur Oxidizing Bacteria (SOB) for plant growth promotion
Fourteen heterorophic sulphur oxidizing bacteria were identified and two isolates S 14 and Ca7 could release 6 µg SO4-/ mg of sulphur. Inoculation with SS1 (Bacillus flexus) for chickpea and BS104 (Alcaligens sp.) for mustard significantly improved the growth and yield in pot and field experiment. Alcaligens sp. BS104, a facultative autotroph identified as the most potent S-oxidizing bacteria in two years field experiment on mustard.
Table: Effect of Sulfur oxidizers on yield and oil content
| Treatments | No. of siliqua/plant | Seed yield (Kg/12 m2) | Oil Content (%) |
|---|---|---|---|
| T1 -NPK | 407.33e | 1.55e | 40.15e |
| T2 NPKS | 410.00e | 1.61bcd | 40.86d |
| T3 BS104 + NPK | 505.00b | 1.94b | 41.41b |
| T4 BS104 +NPKS | 547.00a | 2.25a | 41.78a |
| T5 NPK + Gypsum | 568.67a | 2.39a | 41.72a |
| SEm+ | 7.66 | 0.08 | 0.04 |
| CD | 22.95 | 0.23 | 0.12 |
3. Dissimilatory nitrate reduction to ammonium (DNRA): a means to conserve nitrogen under anaerobic conditions
A simple method was developed to create microaerophilic condition and detect Ammonium production. Bacteria with DNRA activity were identified as Achromobacter, Arthrobacter, Bacillus, Bradyrhizobium, Brevibacterium, Enterobacter, Ensifer, Kocuria, Micrococcus, Planococcus, Paenarthrobacter and Rhizobium. These bacteria convert nitrate to nitrite and to ammonia. Thus conserve nitrogen under anaerobic conditions.
4. Boron tolerant strains for improving boron use efficiency

Twenty seven isolates obtained from rhizospheric soil showed growth on media containing boron ranging from 20ppm-80ppm. These cultures were positive for P and Zn solubilization; production of siderophore, ammonia and IAA. Plant studies are in progress to look for B fortification using these microorganisms
5. Biofortification of wheat with Fe and Zn using bacterial endophytes
Inoculation of siderophore producing endophytes Arthrobacter sulfonivorans and DS-68 Enterococcus hirae DS-163 increased the iron concentration in wheat grains from 30 to 49 mg kg-1. Endophytes Bacillus subtilis DS-178 and Arthrobacter sp. DS-179 efficient for Zn solubilisation could increase the Zn concentration in wheat grains from 28 to 42 mg kg-1.Inoculation of endophytes up-regulate the Zinc-Iron Protein transporters which contribute to uptake of Fe & Zn in wheat. There was 1 to 4 fold increase in the expression level of transporter genes.
6. Biopriming of rice seeds with cyanobacteria

A biopriming technique was developed for the coating of rice seeds with potential cyanobacterial strains (Plectonema boreanum, Anabaena doliolum, Nostoc commune and an equi-proportional mix-culture of these three strains) and field trials for two consecutive years were conducted on eight rice varieties (PR-118, PR-113, MTU-1010, MTU-7029, HUR-105, PB-1, PB-115 and BPT-5204). Viability of the coated seeds and cyanobacteria were found significant even after 18 months of coating. Germination of cyano-coated seeds was 18-20% higher than non-coated seeds. Results on agronomic parameters and biochemical tests were encouraging and coating of seeds with cyanobacteria led to an increase of 5.3 to 7.9% increase in grain yield among different varieties.
7. Plant growth promoting bacteria for vegetable crops

A bacterial formulation BIOGROW was developed using a consortium of five bacteria endowed with phosphorus solubilization, IAA and siderophore production attributes. The formulations were tested on tomato for two consecutive years and 25-30% increase in yield was recorded in treatment inoculated with BIOGROWI. Moreover there was a significant improvement in nutritional quality as evident from enhanced content of lycopene and β-carotene in inoculated treatment as compared to control.
ix. Microbe based technologies for biotic stress management
1. Trichoderma and Pseudomonas based formulations for biocontrol
Three bioformulations of P. fluorescens, T. harzianum and T. viride namely Eco-Pesticide (Talc based bioformulation of Pseudomonas fluorescens); Eco-Green Fungicide (Vermi-based bioformulation of T. viride) and Green Fungicide (Talc based bioformulation of Trichoderma harzianum) respectively, were developed successfully and found effective against a number of soil and seed borne pathogens like Rhizoctonia, Sclerotium, Sclerotinia, Fusarium, Pythium, Ralstonia, Macrophomina, Bipolaris, Phoma, etc.

Green fungicide (Talc based formulation of Trichoderma harzianum) tested in papaya and tomato nursery at farmers’ field in Begusarai, Bihar. No seedling mortality from soil-borne pathogens like Pythium, Phytophthora, Rhizoctonia, etc. as compared to pesticides treatments where 40% mortality was recorded.
2. Bio Pulse: A fly ash based formulation for biocontrol of Fusarium wilt in pulses
A bioformulation of Trichoderma harzianum and Bacillus amyloliquefaciens for control of Fusarium wilt in chickpea. Treatment with formulation could suppress wilt disease by 40% and increased the grain yield by 15% in chickpea on farmers’ field.

3. Biofilmed inoculants: a novel strategy for management of collar rot and wilt disease of chickpea

Biofilms were prepared with Trichoderma viride as the matrix and Bacillus subtilis as partner.A synthetic medium named Z-medium was found to be the most suitable medium in terms of population growth, fresh mass, dry biomass of Trichoderma viride and Bacillus subtilis. Efficacy of biofilm against collar rot and wilt of chickpea was demonstrated in sick pots under net house.
4. Actinomycetes for antagonism against Fusarium oxysporum f. sp. lycopersici, Rhizoctonia solani and Sclerotium rolfsii
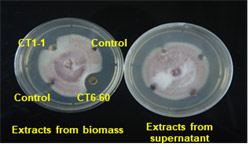
A total of 208 actinomycete isolates were obtained and screened for antagonistic properties against Fusarium oxysporum f. sp. lycopersici, Rhizoctonia solani and Sclerotium rolfsii Interestingly it was observed that the fungal growth was inhibited by mycelial crude extracts but not by supernatantsThe extracts were characterized by LC-MS for identification of active metabolite(s) responsible for inhibition.
5. Endophytes for biocontrol of Sclerotium rolfsii

In planta trial in tomato challenged with pathogen S. rolfsii in presence and absence of endophyte inoculation revealed that Bacillus sp. 2P2 showed the highest protection against S. rolfsii. These strains elicited induced systemic resistance of plant and significantly higher activity (p≤0.05) of phenylalanine ammonia lyase, peroxidase, polyphenol oxidase, and ascorbate oxidase indicating the further strengthening of cell wall barrier through lipid peroxidation, cross linking of cell walls, lignifications, suberization and other cell wall strengthening processes. It was further confirmed by confocal scanning laser micrographs of upper collar region. It was evident that the inoculation of endophyte inhibited the colonization and movement of the pathogen. In addition, endophytes upregulated the expression of three pathogenesis-related genes PR1a, PR2a, and PR3, which are responsible for production of glucanases and chitinases contributing to pathogen inhibition
6. Development of Trichoderma based liquid formulation
A month old Trichoderma was used to filter spores.Centrifugation technique was used to harvest spores without mycelium contamination. To develop spore based liquid formulation of Trichoderma, spore were harvested & mixed with four different concentrations of solvent solution (A). Four months after preparation of formulation, the spores were viable.The decline in CFU count in all treatments (T1 to T4) is rapid as compared to control.
x. Microbe based technologies for abiotic stress management
1. Archaeal formulation for alleviation of drought stress

IThe Bureau has developed an archeal bioformulation which had been used to grow mustard without any irrigation and wheat with one irrigation. Although yield decreased significantly, but the application of bioformulation could supplement 25-30% yield as compared to un-inoculated control under water stress. Such a formulation can immensely help the farmers to harvest some yield instead of nothing under severe drought conditions. This formulation is being validated throughout the country under AICRP-NSP
2. Evaluation of drought alleviating microbes and unravelling the plant-microbe interaction
Molecular plant-microbe interaction has been deciphered for microbe mediated drought alleviation in rice. Drought responsive genes for plasma membrane intrinsic protein (OsPIP), dehydrin, DREB, dehydration responsive element binding, CuZn-SOD, Fe-SOD, Mn-SOD, PAL, Chl_sAPX, CAT, AU076282, D14481 and actin were found to be differentialy expressed in Trichoderma harzianum, Pseudomonas fluorescens inoculated rice plants grown under water stress.

Effect of seed treatment with A. Trichoderma + Pseudomonas, B. Pseudomonas; C. Trichoderma and D. Control (no treatment) on rice
3. Alleviation of heavy metal stress using microorganisms

Ochrobactrum intermedum BB-12 isolated and characterized from metal polluted soils could alleviate the stress caused by high concentration of cadmium on spinach. It could also reduce the uptake of Cd from soil to root to shoot. Inoculation of Bacillus mycoides NR-5 isoalted from arsenic contaminated soil, showed early germination, increase in shoot and root fresh weight to a the tune of19.71 and 36.84% respectively in spinach grown in arsenic spiked soils
4. Alleviation of salinity stress in tomato using endophytes
Endophytes viz. Bacillus safensis BTL5 and Bacillus hayansii GTR8 isolated from Tulsi were found to be highly effective to alleviate salinity stress in tomato. The isolates could improve the root architecture of tomato and also reduced the oxidative stress under saline conditions.
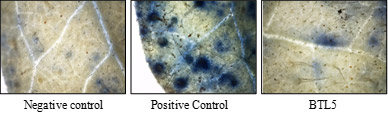
Reduction in superoxide levels in tomato leaves due to inoculation of endophytes.
Significant advancement in science/technology
BGA conservation techniques
Developed procedures for storage of cyanobacteria on filter paper strips and silica gel (mesh size 60-120). With filter paper strips, culture was viable for 12 months as evident from chlorophyll content used as a measure of growth. Using silica gels, cyanobacterial cultures were viable till 18 months.

Phytosterol for control of Alternaria arborescens
Computational studies coupled with wet lab identified phytosterols as a potential inhibitor of growth for Alternaria arborescens. Phytosterols could block the FMN binding sites of chorismate synthase which in turn stops multiple essential metabolic processes.

Purification of phycobilins from cyanobacteria
Following extraction of phycobiliproteins through freeze-thaw and purification through ammonium sulphate precipitation & anion exchange chromatography, phycocyanin (purity= 4.0) and phycoerythrin (purity: 6.0) were purified. The purified pigments were pharmaceutical grade and can be used for multiple biotechnological applications.

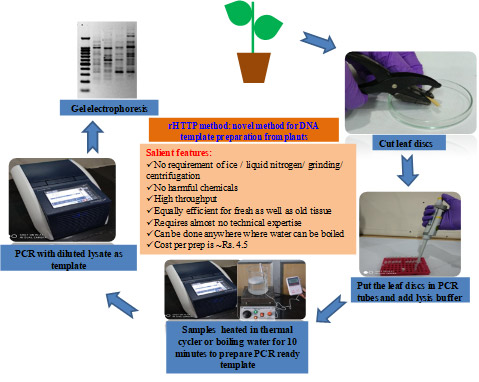
Rapid high throughput template preparation method
A novel high throughput method for preparation of PCR ready templates from a wide range of plants has been developed. Following this method, PCR ready templates can be prepared in field conditions without any instrumentation, with basic facilities to boil water. This methodology can be used for molecular detection of pathogens.
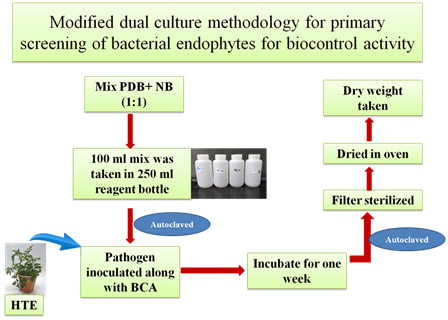
Modified dual culture methodology
In modified methodology of liquid dual culture assay, endophyte bacteria and pathogen are inoculated in bottles broth containing 1:1 ratio of PDB and nutrient broth with sufficient head space and allowed to interact actively for 7 days in the presence of host tissue extract. At the end, inhibition was measured based on biomass accumulated by fungus by the collective effect of any of the mechanisms endophyte might be having. Performing this method for first screening could reduce the chances of losing a potent endophyte from primary population.
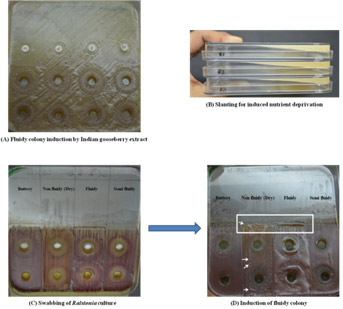
Nutritional and biotic factors for resurgence of virulence in Ralstonia
Major problem faced with this pathogen is rapid loss of virulence in lab conditions; therefore, the method has been devised to induce virulence. The method precisely uses physical and chemical stimulators for maintaining virulent cultures of Ralstonia solanacearum. The method has been successfully tested on different strains of Ralstonia.

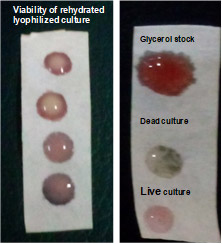
Strip for viability testing of bacteria
A paper based strip has been developed for testing viability of the bacteria without any requirement to cultivate them. This strips can be used for testing of viability of bacteria from glycerol stocks, lyophilized vials or formulations.
Microscope assisted slide –dilution method for isolation and purification of bacteria
A new, easy and effective method for isolation and purification of cyanobacteria has been developed. This method was able to isolate and purify cyanobacteria which were otherwise difficult to isolate using routine repeated subculturing methods

Method for high throughput template preparation from fungi
A method for rapid PCR ready template preparation from fungi has been developed. It is high throughput, cost effective, requires very minimal instrumentation and easy to perform. The templates prepared following this method can be used for PCr amplification and subsequent downstream applications.
MicroVeda
A comprehensive web based system for agriculturally important microbes namely www.microveda.org.inis an unique database having information on the collection of microbes important to agriculture and allied sectors in India, their geo location, characteristics and importance, etc. Also the database attempts to incorporate the Location based searching; MIicrobialDAtabase Search (MIDAS); Knowledge models of microorganisms and trait wise listing of elite microbes at NAIMCC. The database was released by Dr. A.N. Mukhopadhyay, Chairman RAC on 15th February, 2019 during RAC meeting
Key highlights of the database is
- Unique database defining geo-locations of isolated Agriculturally Important Microorganisms (AIM) strains
- Comprehensive information about AIM w.r.t. characteristics and importance
- Knowledge models and Microbial Database Search (MIDAS) giving concise information
- Trait wise listing of elite microbes at NAIMCC
Chitin-fortified Trichoderma formulation for the management of tomato root rot caused by Rhizoctonia solani
Chitin-fortified Trichoderma formulation offered cost-effective and improved biocontrol of tomato root rot and wilt caused by Rhizoctonia solani and methodology mentioned for formulation development is used by several researchers as baseline for their biocontrol studies
Sequencing of metgenomes and metaproteomes
Shotgun sequencing of various niches like Khardoong La Subglacial Soil, Pangong Lake, Leh,Mangrove soil, Saline soil, Solid waste, Wheat rhizosphere from different regions as well as saline soil, Arsenic contaminated soil; Valley of Flowers and Termite gut have been done and information on composition and functions have been derived from multitudes of uncultivable prokaryotic and eukaryotic species and complex microbial population existing in particular environment. Bureau has also taken lead in standardizing protocol for recovering all proteins recovered directly from environmental samples at a given time i.e. metaproteome and first time reference set of proteomic data of microbial communities associated with maize rhizosphere have been made available. Subsequently metaproteome of wheat rhizosphere from saline and non saline region has also been compared.
Whole genome sequencing
Bureau has been continuously involved in deciphering capabilities of microbes by genomic analysis tools and techniques. ICAR-NBAIM sequenced country’s first complete draft genome of Mesorhizobium ciceri strainCa181 in 2011.Whole genome sequencing of various agriculturally important microbes Exiguobacterium profundum PHM 11, Bacillus subtilis RC 25, Pseudomonas azotoformans SC 14, Chromohalobacter salexigens ANJ 207, Pseudomonas koreensis P2, Staphylococcus xylosusLSR_02N, Brevibacillus borstelensis LCHU R05have been done and information on stress responsive traits, metabolic pathways and transporter profiles have been obtained. The bureau has published the first draft genome sequence of Fusarium udum.

